One Row to Multiple Rows & Dynamic Columns
【Question】
I have a CSV file whose awful format I cannot change (simplified here):
Inc,a_One,a_Two,a_Three,b_One,b_Two,b_Three
1,1,1.5,"5 Things",2,2.5,"10 Things"
2,5,5.5,"10 Things",6,6.5,"20 Things"
Inc,a_One,a_Two,a_Three,b_One,b_Two,b_Three
3,9,9.5,"15 Things",10,10.5,"30 Things"
My desired output is a new CSV containing:
inc,label,one,two,three
1,"a",1,1.5,"5 Things"
2,"a",5,5.5,"10 Things"
3,"a",9,9.5,"15 Things"
1,"b",2,2.5,"10 Things"
2,"b",6,6.5,"20 Things"
3,"b",10,10.5,"30 Things"
Basically:
· lowercase the headers
· strip off header prefixes and preserve them by adding them to a new column
· remove header repetitions in later rows
· stack each column that shares the latter part of their names (e.g. a_One and b_One values should be merged into the same column).
· During this process, preserve the Inc value from the original row (there may be more than one row like this in various places).
With caveats:
· I don't know the column names ahead of time (many files, many different columns). These need to be parsed if they are to be used as logic for stripping the repetitious header rows.
· There may or may not be more than one column with properties like Inc that need to be preserved when everything gets stacked. Generally, Inc represents any column that does not have a prefix like a_ or b_. I have a regex to strip out these prefixes already.
So far, I've accomplished this:
> wip_path <- 'C:/path/to/horrible.csv'
> rawwip <- read.csv(wip_path, header = FALSE, fill = FALSE)
> rawwip
V1 V2 V3 V4 V5 V6 V7
1 Inc a_One a_Two a_Three b_One b_Two b_Three
2 1 1 1.5 5 Things 2 2.5 10 Things
3 2 5 5.5 10 Things 6 6.5 20 Things
4 Inc a_One a_Two a_Three b_One b_Two b_Three
5 3 9 9.5 15 Things 10 10.5 30 Things
> skips <- which(rawwip$V1==rawwip[1,1])
> skips
[1] 1 4
> filwip <- rawwip[-skips,]
> filwip
V1 V2 V3 V4 V5 V6 V7
2 1 1 1.5 5 Things 2 2.5 10 Things
3 2 5 5.5 10 Things 6 6.5 20 Things
5 3 9 9.5 15 Things 10 10.5 30 Things
> rawwip[1,]
V1 V2 V3 V4 V5 V6 V7
1 Inc a_One a_Two a_Three b_One b_Two b_Three
But then when I try to apply a tolower() to these strings, I get:
> tolower(rawwip[1,])
[1] "4" "4" "4" "4" "4" "4" "4"
And this is quite unexpected.
So my questions are:
1)How can I gain access to the header strings in `rawwip[1,]` so that I can reformat them with `tolower()` and other string-manipulating functions?
2) Once I’ve done that, what’s the most effective way to stack the columns with shared names while preserving the inc value for each row?
Bear in mind, there will be well over a thousand repetitious columns that can be filtered down to perhaps 20 shared column names. I will not know the position of each stackable column ahead of time. This needs to be determined within the script.
【Answer】
It’s easy to implement a static transposition in R, but it’s hard to perform a dynamic one in the language. A dynamic transposition is like this: field names can be any strings (not limited to a, b, one, two, three) and the number of field names is unlimited (not limited to 2*3). It’s convenient to achieve the dynamic transposition in SPL. Here’s the SPL script:
| A |
B |
|
| 1 |
=file("d:\\source.csv").import@tc() |
|
| 2 |
=A1.select(Inc!="Inc") |
|
| 3 |
=A1.fname().to(2,).(~.split("_")) |
|
| 4 |
=A3.id(~(1)) |
=A3.id(~(2)) |
| 5 |
=create(inc,label,${B4.(lower(~)).concat@c()}) |
|
| 6 |
=A2.run(A4.run(A5.record(A2.Inc|A4.~|B4.(A2.~.field(A4.~+"_"+~))))) |
|
| 7 |
=file("d:\\result.csv").export@tc(A5) |
|
A1: Import the CSV file and return it as a table sequence;
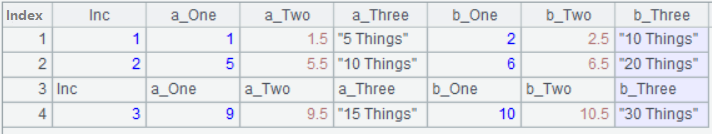
A2: Select data that doesn’t include the headers;

A3: Split filed names that contain "_"; the result is a sequence of sequences;
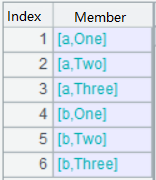
A4: Remove duplicates;

B4: Remove duplicates;

A5: Create a new table sequence to store the final result; macro replacement is used here to lowercase B4’s strings;
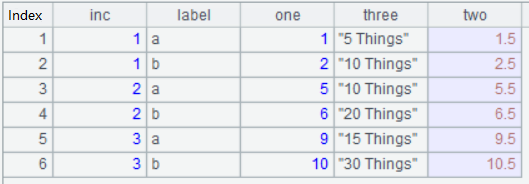
A6: Transpose each of A2’s records into multiple records and store the result in A5’s table sequence; the above pic is the transposed table;
A7: Export A5’s result to a new CSV.
SPL Official Website 👉 http://www.scudata.com
SPL Feedback and Help 👉 https://www.reddit.com/r/esProc
SPL Learning Material 👉 http://c.scudata.com
SPL Source Code and Package 👉 https://github.com/SPLWare/esProc
Discord 👉 https://discord.gg/ydhVnFH9
Youtube 👉 https://www.youtube.com/@esProc_SPL


