How Python Handles Big Files
The Python programming language has become more and more popular in handling data analysis and processing because of its certain unique advantages. It’s easy to read and maintain. pandas, with a rich library of functions and methods packaged in it, is a fast, flexible and easy to use data analysis and manipulation tool built on top of Python. It is one of the big boosters to make Python an efficient and powerful data analysis environment.
pandas is memory-based. It does a great job when the to-be-manipulated data can fit into the memory. It is inconvenient, even unable, to deal with big data, which can’t be wholly loaded into the memory. Large files, however, like those containing data imported from the database or downloaded from the web, are common in real-world businesses. We need to have ways to manage them. How? That’s what I’d like to say something about.
By “big data” here, I am not talking about the TB or PB level data that requires distributed processing. I mean the GB level file data that can’t fit into the normal PC memory but can be held on disk. This is the more common type of big file processing scenario.
Since a big file can’t be loaded into the memory at once, we often need to retrieve it line by line or chunk by chunk for further processing. Both Python and pandas support this way of retrieval, but they don’t have cursors. Because of the absence of a cursor mechanism, we need to write code to implement the chunk-by-chunk retrieval in order to use it in functions and methods; sometimes we even have to write code to implement functions and methods. Here I list the typical scenarios of big file processing and their code examples to make you better understand Python’s way of dealing with them.
I. Aggregation
A simple aggregation is to traverse values in the target column and to perform calculation according to the specified aggregate operation, such as the sum operation that adds up traversed values; the count operation that records the number of traversed values; and the mean operation that adds up and counts the traversed values and then divides the sum by the number. Here let’s look at how Python does a sum.
Below is a part of a file:

To calculate the total sales amount, that is, doing sum over the amount column:
1. Retrieve file line by line
total=0 with open("orders.txt",'r') as f: line=f.readline() while True: line = f.readline() if not line: break total += float(line.split("\t")[4]) print(total) |
Open the file Read the header row
Read detail data line by line Reading finishes when all lines are traversed
Get cumulated value |
2. Retrieve file chunk by chunk in pandas
pandas supports data retrieval chunk by chunk. Below is the workflow diagram:
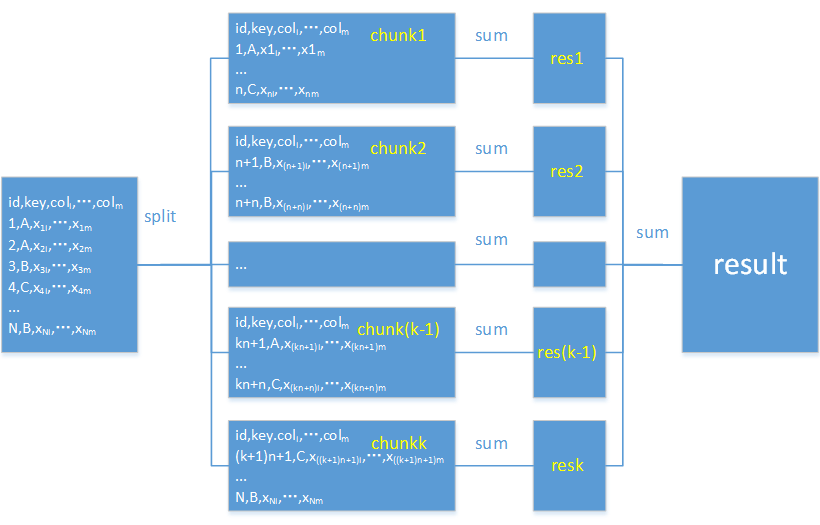
import pandas as pd chunk_data = pd.read_csv("orders.txt",sep="\t",chunksize=100000) total=0 for chunk in chunk_data: total+=chunk['amount'].sum() print(total) |
Retrieve the file chunk by chunk; each contains 100,000 lines
Add up amounts of all chunks
|
Pandas is good at retrieval and processing in large chunks. In theory, the bigger the chunk size, the faster the processing. Note that the chunk size should be able to fit into the available memory. If the chunksize is set as 1, it is a line-by-line retrieval, which is extremely slow. So I do not recommend a line-by-line retrieval when handling large files in pandas.
II. Filtering
The workflow diagram for filtering in pandas:
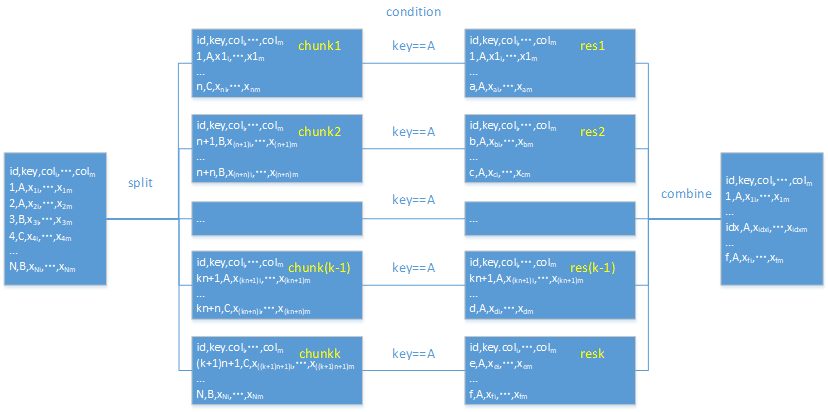
Similar to the aggregation, pandas will divide a big file into multiple chunks (n), filter each data chunk and concatenate the filtering results.
To get the sales records in New York state according to the above file:
1. With small data sets
import pandas as pd chunk_data = pd.read_csv("orders.txt",sep="\t",chunksize=100000) chunk_list = []
for chunk in chunk_data: chunk_list.append(chunk[chunk.state=="New York"]) res = pd.concat(chunk_list) print(res) |
Define an empty list for storing the result set
Filter chunk by chunk
Concatenate filtering results |
2. With big data sets
import pandas as pd chunk_data = pd.read_csv("orders.txt",sep="\t",chunksize=100000) n=0 for chunk in chunk_data: need_data = chunk[chunk.state=='New York'] if n == 0: need_data.to_csv("orders_filter.txt",index=None) n+=1 else: need_data.to_csv("orders_filter.txt",index=None,mode='a',header=None) |
For the result set of processing the first chunk, write it to the target file with headers retained and index removed
For the result sets of processing other chunks, append them to the target file with both headers and index removed |
The logic of doing aggregates and filters is simple. But as Python doesn’t provide the cursor data type, we need to write a lot of code to get them done.
III. Sorting
The workflow diagram for sorting in pandas:
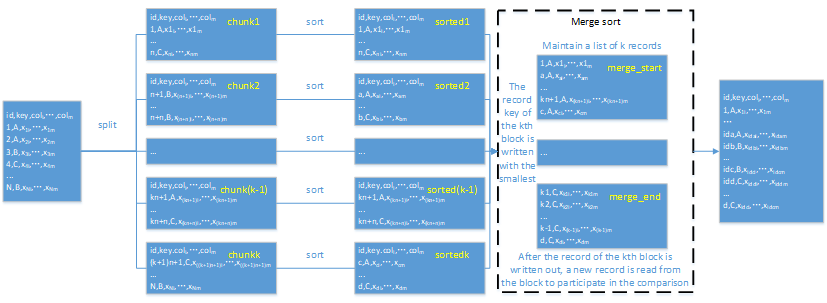
Sorting is complicated because you need to:
1. Retrieve one chunk each time;
2. Sort this chunk;
3. Write the sorting result of each chunk to a temporary file;
4. Maintain a list of k elements (k is the number of chunks) into which a row of data in each temporary file is put;
5. Sort records in the list by the sorting field (same as the sort direction in step 2);
6. Write the record with smallest (in ascending order) or largest (in descending order) value to the result file;
7. Put another row from each temporary file to the list;
8. Repeat step 6, 7 until all records are written to the result file.
To sort the above file by amount in ascending order, I write a complete Python program of implementing the external sorting algorithm:
import pandas as pd import os import time import shutil import uuid import traceback
def parse_type(s): if s.isdigit(): return int(s) try: res = float(s) return res except: return s
def pos_by(by,head,sep): by_num = 0 for col in head.split(sep): if col.strip()==by: break else: by_num+=1 return by_num
def merge_sort(directory,ofile,by,ascending=True,sep=","):
with open(ofile,'w') as outfile:
file_list = os.listdir(directory)
file_chunk = [open(directory+"/"+file,'r') for file in file_list] k_row = [file_chunk[i].readline()for i in range(len(file_chunk))] by = pos_by(by,k_row[0],sep)
outfile.write(k_row[0]) k_row = [file_chunk[i].readline()for i in range(len(file_chunk))] k_by = [parse_type(k_row[i].split(sep)[by].strip()) for i in range(len(file_chunk))]
with open(ofile,'a') as outfile:
while True: for i in range(len(k_by)): if i >= len(k_by): break
sorted_k_by = sorted(k_by) if ascending else sorted(k_by,reverse=True) if k_by[i] == sorted_k_by[0]: outfile.write(k_row[i]) k_row[i] = file_chunk[i].readline() if not k_row[i]: file_chunk[i].close() del(file_chunk[i]) del(k_row[i]) del(k_by[i]) else: k_by[i] = parse_type(k_row[i].split(sep)[by].strip()) if len(k_by)==0: break
def external_sort(file_path,by,ofile,tmp_dir,ascending=True,chunksize=50000,sep=',', os.makedirs(tmp_dir,exist_ok=True)
try: data_chunk = pd.read_csv(file_path,sep=sep,usecols=usecols,index_col=index_col,chunksize=chunksize) for chunk in data_chunk: chunk = chunk.sort_values(by,ascending=ascending) chunk.to_csv(tmp_dir+"/"+"chunk"+str(int(time.time()*10**7))+str(uuid.uuid4())+".csv",index=None,sep=sep) merge_sort(tmp_dir,ofile=ofile,by=by,ascending=ascending,sep=sep) except Exception: print(traceback.format_exc()) finally: shutil.rmtree(tmp_dir, ignore_errors=True)
if __name__ == "__main__": infile = "D:/python_question_data/orders.txt" ofile = "D:/python_question_data/extra_sort_res_py.txt" tmp = "D:/python_question_data/tmp" external_sort(infile,'amount',ofile,tmp,ascending=True,chunksize=1000000,sep='\t') |
Function Parse data type for the string
Function Find the position of the column name by which records are ordered in the headers
Function External merge sort
List temporary files
Open a temporary file
Read the headers
Get the position of column name by which records are ordered among the headers Export the headers Read the first line of detail data
Maintain a list of k elements to store k sorting column values
Perform sort in the order of the list Export the row with the smallest value Read and process temporary files one by one
If the file traversal isn’t finished, continue reading and update the list Finish reading the file
Function External sort
Create a directory to store the temporary files
Retrieve the file chunk by chunk
Sort the chunks one by one
Write the sorted file
External merge sort
Delete the temporary directory
Main program
Call the external sort function |
Python handles the external sort using line-by-line merge & write. I didn’t use pandas because it is incredibly slow when doing the line-wise retrieval. Yet it is fast to do the chunk-wise merge in pandas. You can compare their speeds if you want to.
The code is too complicated compared with that for aggregation and filtering. It’s beyond a non-professional programmer’s ability. The second problem is that it is slow to execute.
The third problem is that it is only for standard structured files and single column sorting. If the file doesn’t have a header row, or if there are variable number of separators in rows, or if the sorting column contains values of nonstandard date format, or if there are multiple sorting columns, the code will be more complicated.
IV. Grouping
It’s not easy to group and summarize a big file in Python, too. A convenient way out is to sort the file by the grouping column and then to traverse the ordered file during which neighboring records are put to same group if they have same grouping column values and a record is put to a new group if its grouping column value is different from the previous one. If a result set is too large, we need to write grouping result before the memory lose its hold.
It’s convenient yet slow because a full-text sorting is needed. Generally databases use the hash grouping to increase speed. It’s effective but much more complicated. It’s almost impossible for non-professionals to do that.
So, it’s inconvenient and difficult to handle big files with Python because of the absence of cursor data type and relevant functions. We have to write all the code ourselves and the code is inefficient.
If only there was a language that a non-professional programmer can handle to process large files. Luckily, we have esProc SPL.
It’s convenient and easy to use. Because SPL is designed to process structured data and equipped with a richer library of functions than pandas and the built-in cursor data type. It handles large files concisely, effortlessly and efficiently.
1. Aggregation
A |
|
1 |
=file(file_path).cursor@tc() |
2 |
=A1.total(sum(col)) |
2. Filtering
A |
B |
|
1 |
=file(file_path).cursor@tc() |
|
2 |
=A1.select(key==condition) |
|
3 |
=A2.fetch() |
/ Fetch data from a small result set |
4 |
=file(out_file).export@tc(A2) |
/ Write a large result set to a target file |
3. Sorting
A |
|
1 |
=file(file_path).cursor@tc() |
2 |
=A1.sortx(key) |
3 |
=file(out_file).export@tc(A2) |
4. Grouping
A |
B |
|
1 |
=file(file_path).cursor@tc() |
|
2 |
=A1.groups(key;sum(coli):total) |
/ Return a small result set directly |
3 |
=A1.groupx(key;sum(coli):total) |
|
4 |
=file(out_file).export@tc(A3) |
/ Write a large result set to a target file |
SPL also employs the above-mentioned HASH algorithm to effectively increase performance.
SPL has the embedded parallel processing ability to be able to make the most use of the multi-core CPU to boost performance. A @m option only enables a function to perform parallel computing.
A |
|
1 |
=file(file_path).cursor@mtc() |
2 |
=A1.groups(key;sum(coli):total) |
There are a lot of Python-version parallel programs, but none is simple enough.
SPL Official Website 👉 http://www.scudata.com
SPL Feedback and Help 👉 https://www.reddit.com/r/esProc
SPL Learning Material 👉 http://c.scudata.com
SPL Source Code and Package 👉 https://github.com/SPLWare/esProc
Discord 👉 https://discord.gg/ydhVnFH9
Youtube 👉 https://www.youtube.com/@esProc_SPL


